A. Chwialkowska et al. / Bioorg. Med. Chem. Lett. 25 (2015) 3100–3104
3103
RNA substrate (prepared at 0.4–0.8
l
M concentration, reaction vol-
giving in MALDI-TOF MS signals at higher m/z. Recently, we have
reported that in ASL RNA models the guanosine units located in a
single stranded regions of the loop are primary sites for action of
the manganese porphyrin/oxone oxidizing system, resulting in
dehydroguanidinohydantoin and spiroiminodihydantoin deriva-
tives of molecular weight increased by 4 and 32 amu, respec-
umes ca. 1 mL). The reactions were carried out either at room tem-
perature or at 0 °C for 5–180 min, and were terminated by loading
on a C18 Sep-Pack cartridge (Waters) and washing with sterile
water. The RNA products were released from the column by elution
with 50% aq acetonitrile, and quantified spectrophotometrically.
The products (obtained in the yields close to quantitative) were
analyzed by MALDI-TOF mass spectrometry in a negative ions
mode. The effectiveness of the S2U?H2U transformation was cal-
culated from the MALDI-TOF mass spectra, where the intensity of a
band at the m/z attributed to the starting R5S2U-RNA was com-
pared with that at m/z-32. We have earlier demonstrated that
molecular ions for RNA oligonucleotides containing S2U or H2U
are of similar intensities, what indicates their similar ionization
1
5
tively. Also other guanosine lesions having higher M.W. than
1
6
the starting RNA were reported. Therefore, it is possible that gua-
nosine residues in oligomers 3 and 9 might get oxidized, although
detailed explanation cannot be offered, especially because in the
remaining examples we have observed only the expected
S2U?H2U desulfuration. Selected mass spectra recorded after
desulfuration of 9 over different time and oxidant concentration
conditions are presented in Figure S14 (annotations m5H2U-
7
ox
properties. The product of desulfuration of oligomer 1 was posi-
RNA indicate the unidentified oxidized products).
tively identified by IE-HPLC co-injection with corresponding syn-
To check whether H2U product can be obtained from S2U
located in the double-stranded oligonucleotide, 1 was hybridized
thetically prepared H2U-RNA.7
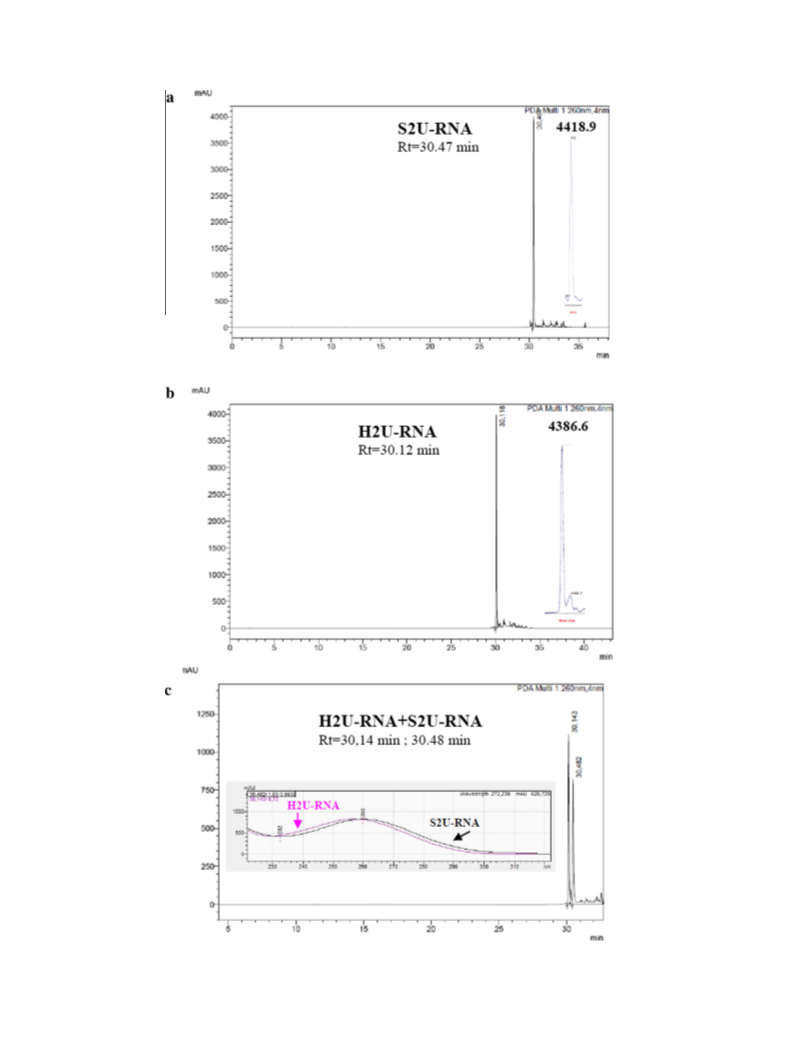
The example of the desulfuration of S2U-RNA 1 monitored by
RP-HPLC is shown at Figure 2. The profile of H2U-RNA clearly indi-
cates that the process of S2U-RNA desulfuration undergoes selec-
tively. Co-injection of both compounds shows that the H2U-RNA
product is less hydrophobic, and its UV spectrum is shifted towards
lower wavelength due to the presence of H2U component
m
with an RNA complement (T = 64.3 °C for the duplex, in 10 mM
1
1
phosphate buffer, 50 mM NaCl) and treated with aqueous oxone
under conditions earlier established for single-stranded 1 (10ꢁ
molar excess of oxone, rt, 5 min.). Standard work-up delivered
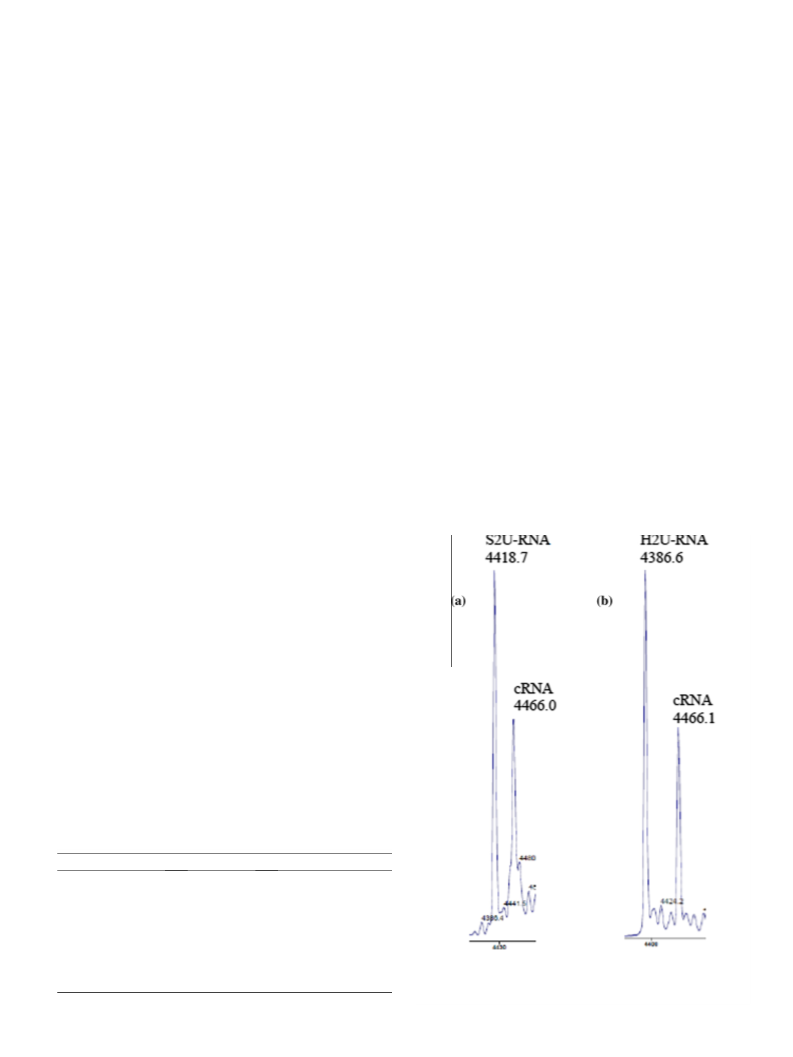
the product identified by MALDI-TOF MS (Fig. 3) as the expected
H2U-RNA/RNA product (m/z 4386.6, M.W. 4392).
(
k
max = 242 nm, see Fig. S2).
The H2U-RNA product was also digested with nuclease P1 and
In conclusion, we found that several 2-thiouridine nucleosides
present in an RNA chain (in single stranded and double stranded
models), can be post-synthetically transformed to corresponding
desulfured 4-pyrimidinone riboside products. The reactions effi-
ciently undergo with 10-fold molar excess of aqueous oxone, at
room temperature, but the reaction time has to be controlled and
optimized. Longer exposition of the R5H2U-RNA products to the
alkaline phosphatase (Sigma–Aldrich), and the resultant mixture
of nucleosides was analyzed by RP-HPLC. The identity of the
released H2U nucleoside was confirmed by co-injection with the
genuine H2U nucleoside, and by UV analysis (see Figs. S1–S3,
Supporting information).10
The results indicate that low excess of oxone is beneficial for
yield and purity of the desired desulfured R5H2U-RNAs (Table 2).
For the oxidation with a 10-fold molar excess of oxone at room
temperature, the reaction time was optimized in the range of 5–
1
20 min (Table 2). The mass spectra for substrates 1–10 and for
crude mixtures containing their R5H2U-RNA desulfured products
are shown in the Supplementary material (Fig. S4–S13). In the
spectra, the bands expected for R5H2U-RNA were accompanied
by those at m/z+22, +38 or +60, which can be assigned to ions
+
+
+
+
which acquired Na , K , or Na and K cations, respectively. It
was found that the reactions lasting for not more than 30 min
furnished the expected products in the yields >90%, except for 5
containing mnm5S2U located in the loop region (87% yield), for
which certain amount of uridine-RNA was observed. Possible
residual amounts of R5U-RNA products obtained during the desul-
furation reaction or R5S2U-RNA contaminations introduced into
the reaction mixture with the R5S2U substrate exhibit signals of
m/z bigger by 16 amu comparing R5H2U-RNA signals.
All R5H2U-RNA products but these obtained from m5S2U (9)
and
sm5S2U (3) were stable in the oxidation conditions for
1
80 min. The two less stable were prone for further oxidation,
Table 2
Optimized time and yield of R5S2U-RNA transformation to R5H2U-RNA for the
reaction carried out with 10-fold molar excess of oxone, at 25 °C
No. of oligomer
Time (min)
Yield of the R5H2U-RNA (%)
1
2
3
4
5
6
7
8
9
10
10
5
30
10
30
10
10
5
>95
93
>95
>95
87
>95
94
>95
>95
>95
a
Figure 3. MALDI-TOF mass spectrometry analysis: (a) S2U-RNA 1 (m/z 4418.7, MW
0
0
4423) hybridized with the RNA complement 3 -GCCGAAAAUUGGCU-5 , (cRNA; m/z
466.0, M.W. 4470); (b) the product of desulfuration, H2U-RNA (m/z 4386.6) and
1
0
10
4
a
1
5-Fold excess.
the unchanged RNA complement (m/z 4466.1).

 Chwialkowska, Anna
Chwialkowska, Anna
 Wielgus, Ewelina
Wielgus, Ewelina
 Leszczynska, Grazyna
Leszczynska, Grazyna
 Sobczak, Milena
Sobczak, Milena
 Mikolajczyk, Barbara
Mikolajczyk, Barbara
 Sochacka, Elzbieta
Sochacka, Elzbieta
 Nawrot, Barbara
Nawrot, Barbara