Angewandte
Research Articles
Chemie
À1
1
000 Ug ). Using the same substrate with a succinic semi-
aldehyde dehydrogenase from Escherichia coli resulted in
À1
À1
1
.5% residual activity (2000 Ug instead of 150000 Ug ).
Testing a handful of other aldehyde dehydrogenases mostly
yielded similarly strongly reduced activities, with one excep-
tion: An aldehyde dehydrogenase from Geobacillus stearo-
À1
thermophilus that has nearly identical speed (2000 Ug ) for
the oxidation of butanal to butyric acid (Figure S12). How-
ever, when the less accepted isobutanal is chosen as substrate
+
(
Figure S11), a discrepancy arises and carba-NADP per-
forms three times worse. Somewhat in between are the a-
ketoglutaric semialdehyde dehydrogenase from Pseudomo-
nas putida to run at 40% velocity under the tested conditions
À1
À1
(
7000 Ug instead of 18000 Ug ).
Looking deeper into the kinetic parameters of a selection
of five enzymes shows important features that come along
with the new cofactor (Table S5). The first enzyme we
analyzed is a promiscuous thermostable glucose dehydrogen-
ase originating from the archaeon Saccharolobus solfataricus
Figure 3. A) Reaction scheme for tested reductive aminases AspRedAm
and AdRedAm converting hexanal and various amine sources utilizing
carba-NADPH. B) Activities with reductive aminases using NADPH or
carba-NADPH, hexanal, and various primary amines. Reaction condi-
(
SsGDH). This enzyme was shown to act on a set on
[
36]
nicotinamide biomimetics. The activity of SsGDH against
d-glucose is reduced by approx. 50% with the artificial
cofactor. However, this looks different when the sugars d-
xylose, l-arabinose, and d-galactose are converted (Fig-
ure S3–S6). Substrate inhibition, which occurs in the natural
setup, seems to have completely vanished when carba-
tions: 100 mm Tris-HCl buffer pH 8.2, 5 mm hexanal, 10 mm amine,
À1
1
mm cofactor, 0.2 mgmL enzyme, 308C.
seem to be capable of utilizing a wide range of biomimetic
cofactors. This is not too surprising as the proS-hydride
transfer from the nicotinamide to the flavin can be achieved
efficiently through spatial proximity and does normally
require fewer binding events with buried binding pockets to
stabilize the transition state, meaning that hydride transfer,
and not a structural change, determines the rate in this
+
NADP was chosen as cofactor and specific activities reached
values that were up to six times higher (for d-xylose
À1
À1
12000 Ug instead of 2000 Ug ). We were also able to
detect velocities that were three times higher in combination
with the thermostable PfuADH for reduction and oxidation
reaction at 658C, while BstADH, in contrast, performed
100 times worse in the oxidation of n-butanol, but functioned
[33]
reductive half-reaction.
Besides the FDH mentioned above, further experiments
were performed to test strictly NAD-dependent enzymes
À1
reasonably well in the reduction reaction (1300 Ug instead
[
34]
À1
including diaphorase as well as uronate, borneol, aldehyde,
semialdehyde, glyceraldehyde-3-phosphate, meso-2,3-butane-
diol, glucose and alanine dehydrogenase (Entry 61, 53, 52, 46–
of 1100 Ug ). Another interesting observation is the fact of
a decreased Michaelis constant (K ) for the biomimetic for
m
most of the enzymes, again except for aldehyde dehydrogen-
ases that were analyzed. We have been wondering what could
be the cause of carba-NADP(H) showing such heterogeneous
behavior towards different enzymes, and even more for
different reaction directions in a single enzyme. It appears
that there is no uniform pattern between different enzymes.
Hence, we conclude that the kinetic changes are caused by the
structural difference of both cofactors. In silico calculations
showed a different conformation of the two structures after
energy minimization (cf. in silico part). Bearing in mind that
even small sub-ꢁngstrçm shifts of reaction partners can have
drastic effects on reaction parameters, we think that the
rotation of the nicotinamide and adenine moiety consequent-
ly changes energy levels and thus the binding event during
catalysis. Consequently, different substrates may form altered
interaction patterns with the cofactor. This is also the case for
the BsGDH which shows 50-fold reduced activity with carba-
4
7/55–56, 60, 57, 51, 50, and 58, respectively Table S4). As
expected, we did not find any significant activity in those
reactions.
In general, all enzymes that naturally accept NADP(H)
will also accept carba-NADP(H) with only a few exceptions.
Oxidation of carba-NADPH works extremely well, while
+
reduction of carba-NADP seems to be more difficult in
nearly 30% of all cases. Here activity is reduced by a factor of
2
–10 depending on the enzyme and the substrate. However,
this cannot be generalized since we also found that the
substrate preference changes with the cofactor. It is note-
worthy that the cofactor preference can probably be selec-
tively engineered as different ADH variant activities imply
(
Entry 6–8, Table S4).
An exception to the rule of carba-NADP acceptance is
+
given by many members within the class of aldehyde
+
+
dehydrogenases. This is probably owed to the special NAD-
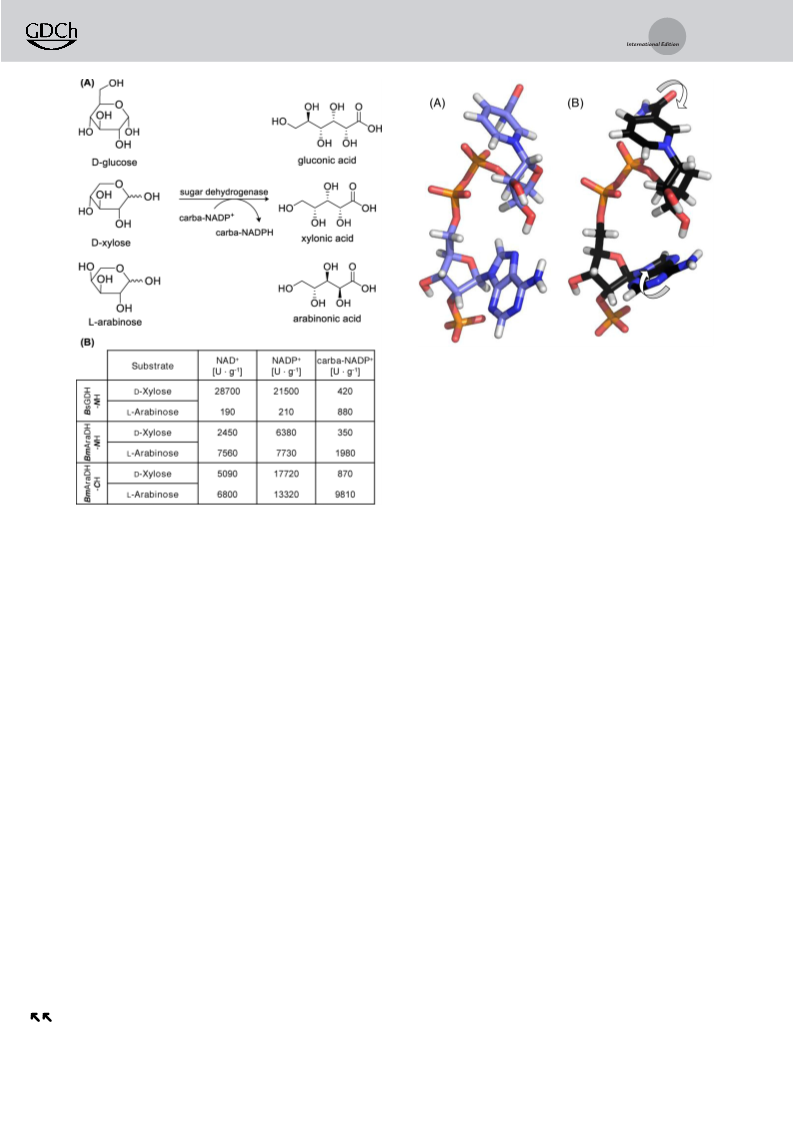
NADP compared to NADP when d-xylose is used as
substrate. However, when using the less accepted substrate l-
arabinose activity increased by a factor of four (Figure 4).
It is well known that affinity-tag additions may result in
+
(
P) binding mode caused by a different Rossman fold type in
[
35]
many aldehyde dehydrogenases. For example, the recently
published aldehyde dehydrogenase from the archaeon Ther-
[37]
moplasma acidophilum showed only 5% residual activity in
altered enzymatic function and protein parameters. Hence,
we also compared one His-tagged dehydrogenase as N-
À1
the oxidation of glyceraldehyde (50 Ug
instead of
Angew. Chem. Int. Ed. 2021, 60, 2 – 8
ꢀ 2021 The Authors. Angewandte Chemie International Edition published by Wiley-VCH GmbH
&&&&
These are not the final page numbers!

 D?ring, Manuel
D?ring, Manuel
 Sieber, Volker
Sieber, Volker
 Simon, Robert C.
Simon, Robert C.
 Tafertshofer, Georg
Tafertshofer, Georg
 Zachos, Ioannis
Zachos, Ioannis