Journal of Agricultural and Food Chemistry
Article
than the wild-type enzyme were selected for rescreening with respect
to expression level and specific activity.
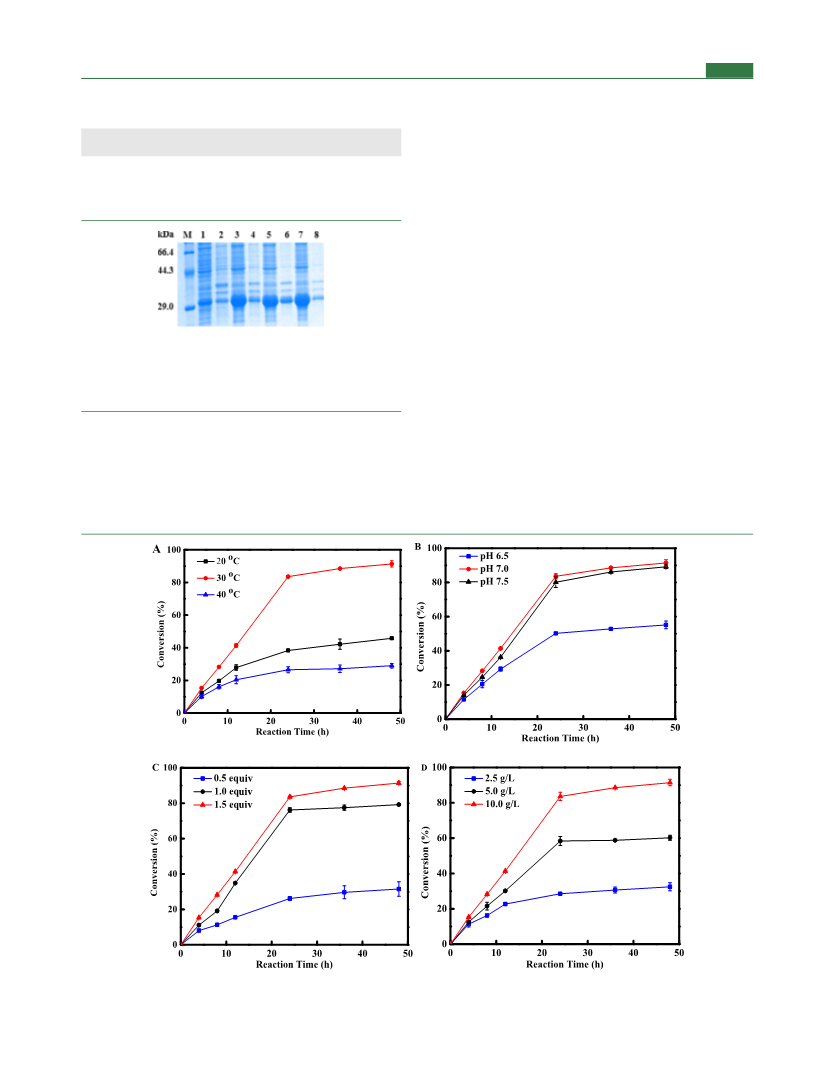
HPLC. The effect of α-KG on the conversion was also investigated at
pH 7.0 and 30 °C with different amounts (0.5, 1.0, and 1.5 equiv to
isoleucine) of α-KG supplemented to the reaction mixture. At last, the
effect of cell loading on the conversion was studied under the
conditions of optimal pH, temperature, and α-KG supplementation.
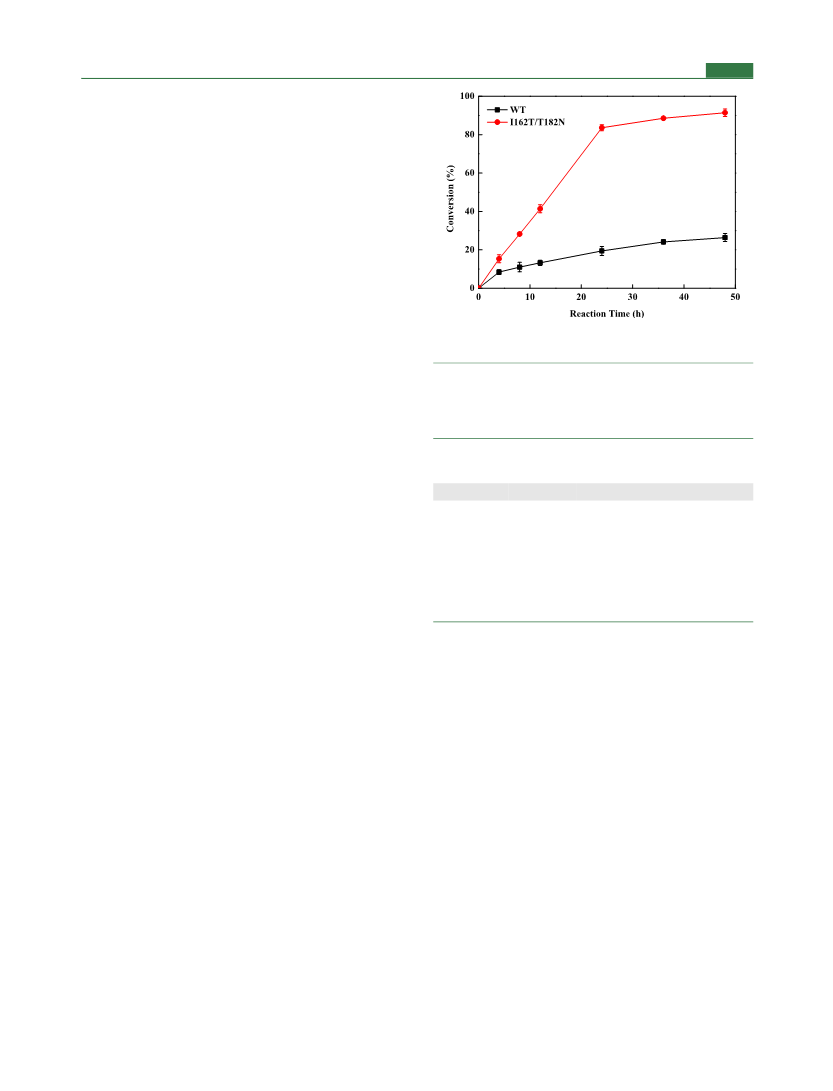
To evaluate the catalytic potential of the variant IDO I162T/
T182N, reaction was carried out on a 50 mL scale with wild-type IDO
as a control. The pH of the reaction mixture composed of 228 mM L-
Ile, 342 mM α-KG, 10.0 mM Fe2+, and 10.0 mM VC was adjusted to
7.0 with 10 N NaOH prior to the addition of 10 g/L of both wild-type
IDO and I162T/T182N in the form of wet cells, and incubation was
carried out at 30 °C and 200 rpm. The amount of (2S,3R,4S)-4-HIL
produced was analyzed by HPLC.
Analytical Methods. L-Ile and (2S,3R,4S)-4-HIL were analyzed
by HPLC with a UV detector at 254 nm and a reverse-phase
Nucleosil C18 column (Hypersil ODS2, 4.6 × 250 mm, 5 μm), and
the column was eluted at a flow rate of 0.8 mL min−1 with a solvent
system of methanol/water (55/45, v/v) at 30 °C.
(2S,3R,4S)-4-HIL isomers were analyzed by HPLC with a UV
detector at 250 nm and a reverse-phase Nucleosil C18 column
(Hypersil ODS2, 4.6 × 250 mm, 5 μm) at 45 °C with a 10 mM, pH
2.8 KH2PO4 (eluent A) and acetonitrile (eluent B) solvent system at a
flow rate of 0.3 mL min−1 in 0−60 min for 20−25% (v/v) B and 25%
(v/v) B in 60.1−70 min.
Expression and Purification of Wild-Type IDO and Its
Mutants. The recombinant E. coli BL21 (DE3) cells were grown at
37 °C and 180 rpm until the OD600 reached 0.6−0.8. IPTG was
added to a final concentration of 0.2 mM, and the cultivation was
continued at 16 °C and 180 rpm for another 24 h. The cells were
harvested, washed twice with saline, and resuspended in ice-chilled
buffer A (25 mM KPB, 300 mM NaCl, 10 mM imidazole, pH 8.0) to
a final concentration of 0.05 g/mL, and then, the cells were disrupted
by sonication. The cell lysate was centrifuged at 4 °C and 8000g for
30 min, and the supernatant was collected as the cell-free extract of
IDO. The supernatant was loaded onto a His-Trap Ni-nitrilotriacetic
acid FF column (5 mL; GE Healthcare Co.) pre-equilibrated with
buffer A. The target protein was eluted using an increasing gradient of
imidazole from 10 to 200 mM at a flow rate of 5 mL/min and
detected by sodium dodecyl sulfate (SDS)-polyacrylamide gel
electrophoresis (PAGE). The fraction containing the purified protein
was collected and concentrated by ultrafiltration. The freshly purified
enzyme was then used for further experiments.
Site-Directed Mutagenesis. Site-directed mutagenesis was
conducted using the recombinant plasmid of pET 28a-IDO as the
template with primers listed in Table S4. PCR amplification
Enzyme Assay. The assay mixture containing 10 mM L-Ile, 10
mM α-KG, 0.5 mM Fe2+, and 0.5 mM Vc in 100 mM KPB buffer (pH
7.0) was incubated with wild-type IDO or its variants at 30 °C for 30
min. The reaction solution was added with an equal volume of
acetonitrile to terminate the enzyme reaction. The resultant solution
was subsequently subjected to derivatization pretreatment, and the
concentration of (2S,3R,4S)-4-HIL was determined by high-perform-
ance liquid chromatography (HPLC). One unit of enzyme activity
was defined as the amount of enzyme required to produce 1 μmol of
(2S,3R,4S)-4-HIL under the assay conditions.
2,3,4,6-Acetyl-β-D-glucopyranosyl isothiocyanate (GITC) precol-
umn derivatization method: the reaction solution was added with an
equal volume of acetonitrile to terminate the reaction, and the
denatured protein was removed by centrifugation. 100 μL of the
supernatant was transferred into a 2 mL Eppendorf tube, to which 150
μL of acetonitrile−water−triethylamine solution (5 mL−5 mL−40
mg) and 250 μL of GITC solution (5 mM, dissolved in acetonitrile)
were added sequentially, and incubated at 30 °C for 30 min.
Characterization of IDO. The optimum temperature for enzyme
activity was determined by performing assays at 20−50 °C and pH
7.0, and the optimum pH was measured by conducting assays at pH
5.0−10.0 at the optimum temperature. The buffers (100 mM) used
were as follows: citric acid-sodium citrate (pH 5.0−6.0), potassium
phosphate (pH 6.0−9.0), and Gly-NaOH (pH 9.0−10.5). The
highest activity was normalized as 100%. To investigate the effect of
metal ions on the activity of IDO, reactions were carried out in 100
mM KPB buffer (pH 7.0) containing 10 mM L-Ile, 10 mM α-KG, 0.5
mM VC, 0.5 mM Fe2+, purified enzyme, and 1 mM of the respective
metal ion [Ca2+, Mg2+, Mn2+, Zn2+, Cu2+, Ni2+, Co2+, Li+, or
ethylenediaminetetraacetic acid (EDTA)] at 30 °C for 5 min.
To investigate the thermostability of IDO, enzyme solutions (1.0
mg/mL) were incubated at different temperatures (30, 40, and 50 °C)
in KPB for different periods, followed by the measurement of the
residual activity. The activity of the enzyme before incubation was
normalized as 100%.
RESULTS AND DISCUSSION
■
Development of a High-Throughput Screening
Method for IDO. A high-throughput screening method
plays a vital role in the identification of improved variants
from a large mutant library,32 especially for the directed
evolution of enzymes without protein structure information
available. In order to develop a high-throughput screening
method for IDO, a dehydrogenase-coupled assay was adopted
(Scheme 2). The activities of the IDO variants were inferred
from the coupled rate of the NAD+ turnover by (2S,3R,4S)-4-
HIL dehydrogenase, which was dependent on the amount of
(2S,3R,4S)-4-HIL formed by IDO. Formation of NADH by
concurrent reduction of NAD+ was detected by the change in
absorbance at 340 nm using a spectrophotometer.33
To verify the linearity of (2S,3R,4S)-4-HIL concentration
with the optical absorbance at 340 nm, different concentrations
of the (2S,3R,4S)-4-HIL commercial sample were added to the
standard reaction mixture and measured by the colorimetric
assay using a microtiter plate reader. A linear relationship
between the absorbance at 340 nm and (2S,3R,4S)-4-HIL
concentration was observed from 0.1 to 0.9 mM (Figure 1A),
which allows for the continuous quantification of the IDO
activity based on the calibration curve. To further validate the
feasibility of this method for screening the mutant library
containing IDO variants with diverse activities, different
amounts of IDO were added to the assay mixture and the
absorbance at 340 nm was measured. Meanwhile, to prevent
possible background interference, a lower concentration of Vc
and Fe2+ (0.05 mM) was adopted. As can be seen from Figure
1B, the absorbance at 340 nm is in good agreement with the
activity of IDO, indicating that this HILDH-coupled assay
method is applicable for the screening of a large library of
IDOs. As compared with previous screening methods, the
method developed in this study is simple, easy-handling, time-
saving, and also suitable for large-capacity random mutation
library screening.
Optimization of Reaction Conditions for IDO-Mediated
Isoleucine Hydroxylation. The reaction conditions for the
enzymatic preparation of (2S,3R,4S)-4-HIL were further optimized
to improve the efficiency. For optimum pH, the reaction mixture (50
mL) containing 228 mM (30 g/L) L-Ile, 342 mM α-KG (1.5 equiv to
isoleucine), 10.0 mM Fe2+, and 10.0 mM VC was adjusted to pH
values of 6.5, 7.0, and 7.5, respectively, prior to the addition of 10 g/L
IDO I162T/T182N wet cells. For optimum temperature, the
reactions were carried out at 20, 30, and 40 °C, respectively, with a
shaker speed of 200 rpm for 48 h, and the amount of the product
(2S,3R,4S)-4-HIL produced in the reaction solution was analyzed by
Directed Evolution of IDO to Improve the Activity
toward L-Ile. Because there is no crystal structure available
for the currently reported IDOs and their homologous
proteins, a structure-guided rational or semirational design
C
J. Agric. Food Chem. XXXX, XXX, XXX−XXX

 Du, Ping
Du, Ping
 Pan, Jiang
Pan, Jiang
 Qian, Xiao-Long
Qian, Xiao-Long
 Xu, Jian-He
Xu, Jian-He
 Yan, Shuai
Yan, Shuai
 Yu, Hui-Lei
Yu, Hui-Lei
 Zhang, Zhi-Jun
Zhang, Zhi-Jun