
Biochemistry p. 4207 - 4217 (2019)
Update date:2022-08-29
Topics:
 Rohweder, Bettina
Rohweder, Bettina
 Lehmann, Gerhard
Lehmann, Gerhard
 Eichner, Norbert
Eichner, Norbert
 Polen, Tino
Polen, Tino
 Rajendran, Chitra
Rajendran, Chitra
 Ruperti, Fabian
Ruperti, Fabian
 Linde, Mona
Linde, Mona
 Treiber, Thomas
Treiber, Thomas
 Jung, Oona
Jung, Oona
 Dettmer, Katja
Dettmer, Katja
 Meister, Gunter
Meister, Gunter
 Bott, Michael
Bott, Michael
 Gronwald, Wolfram
Gronwald, Wolfram
 Sterner, Reinhard
Sterner, Reinhard
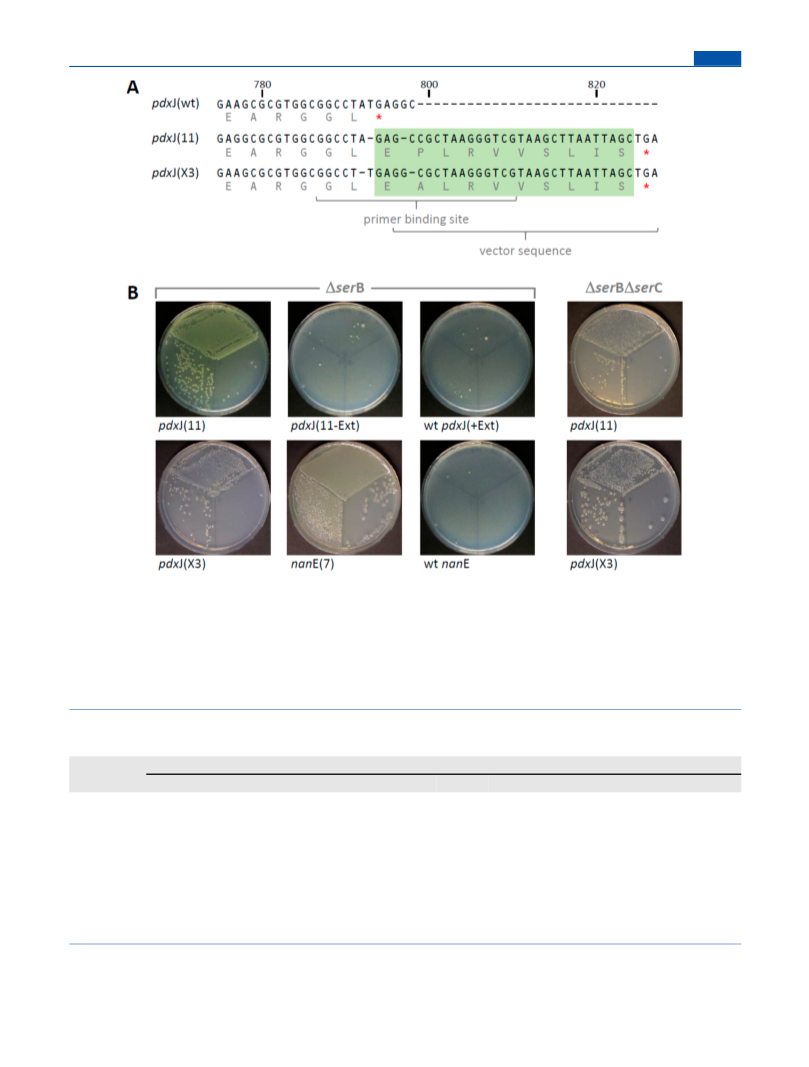
The potential of the frequently encountered (βα)8-barrel fold to acquire new functions was tested by an approach combining random mutagenesis and selection in vivo. For this purpose, the genes encoding 52 different phosphate-binding (βα)8-barrel proteins were subjected to error-prone PCR and cloned into an expression plasmid. The resulting mixed repertoire was used to transform different auxotrophic Escherichia coli strains, each lacking an enzyme with a phosphate-containing substrate. After plating of the different transformants on minimal medium, growth was observed only for two strains, lacking either the gene for the serine phosphatase SerB or the phosphoserine aminotransferase SerC. The same mutants of the E. coli genes nanE (encoding a putative N-acetylmannosamine-6-phosphate 2-epimerase) and pdxJ (encoding the pyridoxine 5′-phosphate synthase) were responsible for rescuing both ΔserB and ΔserC. Unexpectedly, the complementing NanE and PdxJ variants did not catalyze the SerB or SerC reactions in vitro. Instead, RT-qPCR, RNAseq, and transcriptome analysis showed that they rescue the deletions by enlisting the help of endogenous E. coli enzymes HisB and HisC through exclusive up-regulation of histidine operon transcription. While the promiscuous SerB activity of HisB is well-established, our data indicate that HisC is promiscuous for the SerC reaction, as well. The successful rescue of ΔserB and ΔserC through point mutations and recruitment of additional amino acids in NanE and PdxJ provides another example for the adaptability of the (βα)8-barrel fold.
View More









Shijiazhuang Yunxuan Im&Export Co.,Ltd.
Contact:+86-311-83037514
Address:No.6 Hongbin Road
Contact:0510-85393305
Address:1619 Huishan Avenue, Huishan District, Wuxi,
ShanDong XinDa Chemical CO.,LTD
Contact:086-0311-87580543
Address:No.168, High Technology Development Zone Jinan Shandong China
Contact:+91 9963263336
Address:Plot#146A, IDA Mallapur, Hyderabad - 500072
Tai zhou world Pharm & Chem Co., Ltd
Contact:+86-576-85301198
Address:Rome 1001,wangjiang plaza,unti 2,jinshan east Road linhai,zhejiang,china
Doi:10.1017/S1092852900021763
(1957)Doi:10.1038/nature25185
(2018)Doi:10.1007/s11176-005-0336-5
(2005)Doi:10.1016/j.apcata.2015.11.043
(2016)Doi:10.1080/00304940802711275
(2009)Doi:10.1039/c8ra00674a
(2018)