C O M M U N I C A T I O N S
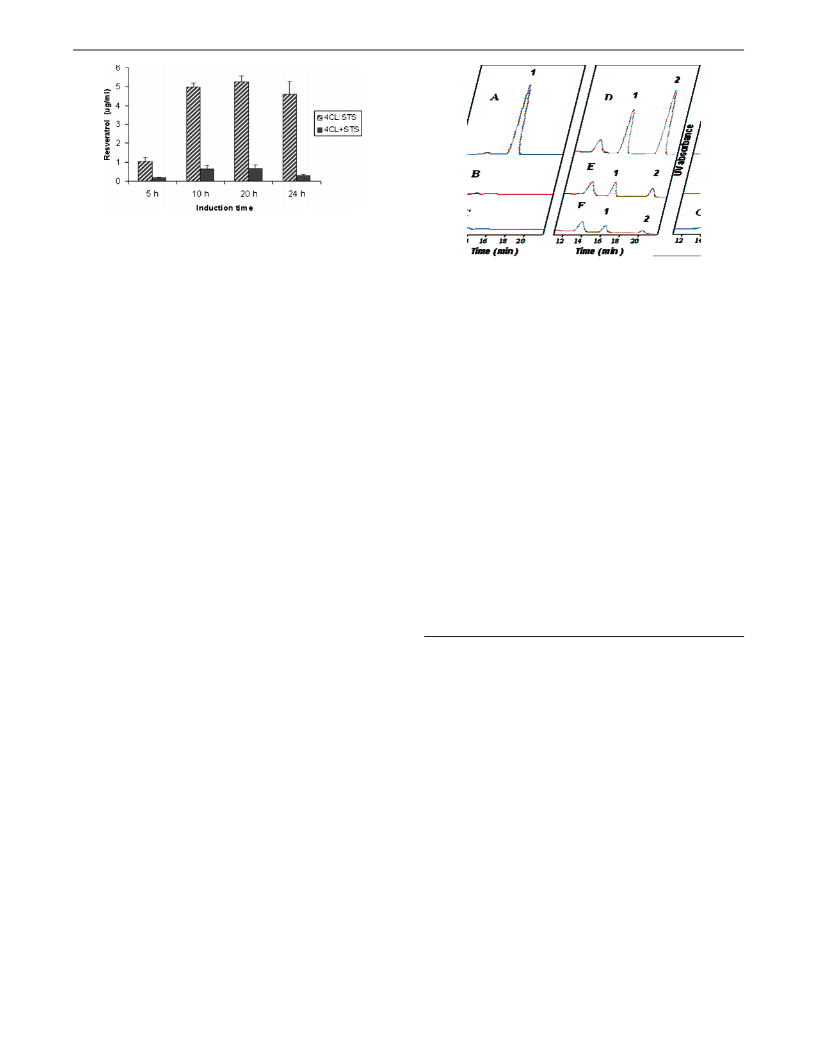
Figure 1. Resveratrol production is lower in the yeast strain cotransformed
with 4CL and STS than in the yeast strain transformed with the 4CL::STS
fusion protein. Error bar is the standard deviation for n ) 3.
production in yeast by up to 15-fold compared to coexpression of
Figure 2. HPLC analysis of control HEK293 cells (A, B, and C) and TAL/
4CL::STS cotransfected HEK293 cells (D, E, and F). A and D were fed
with 4-coumaric acid; B and E were fed with tyrosine; C and F were not
fed with any substrates. Peaks 1 and 2 are 4-coumaric acid and trans-
resveratrol, respectively. All traces were from 290 nm UV profile.
4
CL and STS (Figure 1). Expression of the STS gene was similar
in each yeast strain based on semiquantitative RT-PCR (Figure S3).
After 20 h, yeast expressing the 4CL::STS fusion protein produced
-
1
5
.25 µg mL of resveratrol in the culture, which is 3,500-fold
9
higher than previously reported.
yields by colocalizing 4CL and STS. Moreover, the engineering
of a plant metabolic pathway in mammalian cells may provide
additional opportunities and applications for medical and pharma-
ceutical research. For example, engineered resveratrol by de novo
biosynthesis can conceivably improve cell replacement therapies
by prolonging the life span of donor cells. â-Cell replacement
therapy for type I diabetes using inlet transplantation showed
tremendous promise in recent years, but is somewhat limited by
Because of the difficulty of expressing the TAL gene in yeast
(not shown), we engineered the entire de novo resveratrol biosyn-
thetic pathway into mammalian cells. We cloned the TAL (Figure
S4) into a mammalian transfection vector under the control of a
constitutive CMV promoter. The resulting vector produces a
polycistronic expression unit with TAL and the hygromycin resistant
gene (hpt) linked by an attenuated internal ribosome entry site
(IRES). IRES allows simultaneous translation of TAL and hpt and
13
the short life span of the transplanted â-cells. Although there are
ensures that the hpt gene is transcribed at a lower level than TAL,
leading to high levels of TAL transcription when the transfected
cells are treated with hygromysin.1 Next, we cloned the 4CL::
considerable obstacles to overcome, donor â-cells engineered to
produce resveratrol may survive longer, improving the efficacy and
efficiency of this promising cell replacement therapy.
1
STS fusion gene into another transfection vector that contains a
CMV promoter and a neomycin resistance cassette.12
Acknowledgment. This work was supported by NSF (Grant
MCB0519634), USDA (Grant 2005-05190), and MSMC (Grant 02-
The two constructs were cotransfected into human HEK293
kidney cells. The transfected cells were selected with both G418
and hygromycin for 4 weeks. Both transgenes were highly expressed
in the cotransfected cells based on semiquantitative RT-PCR (Figure
S5). Tyrosine or 4-coumaric acid was added to different batches
of the transfected cells independently, and cultured for 2 days. The
collected cells were resuspended in a potassium phosphate buffer
242) grants to O.Y. and NSF-DBI Grant 0521250 to X.W., J.G.J.,
and O.Y. We thank Peter Murphy for technical support.
Supporting Information Available: Experimental details and
additional results. This material is available free of charge via the
Internet at http://pubs.acs.org.
(pH 7.0), sonicated, and extracted with ethyl acetate. The extracts
References
were analyzed by HPLC and LC-MS. When fed with 4-coumaric
acid (12 µg mL ), the cotransfected lines showed a resveratrol
peak not found in the control at the correct retention time (Figure
(
1) Winkel, B. S. J. Annu. ReV. Plant Physiol. Mol. Biol. 2004, 55, 85-107.
-
1
(2) Schr o¨ der, J.; Schr o¨ der, G. Z. Naturforsch. 1990, 45, 1-8.
(
3) Jang, M.; Cai, L.; Udeani, G. O.; Slowing, K. V.; Thomas, C. F.; Beecher,
C. W.; Fong, H. H.; Farnsworth, N. R.; Kinghorn, A. D.; Mehta, R. G.;
Moon, R. C.; Pezzuto, J. M. Science 1997, 275, 218-220.
2A, D) with an identical UV spectrum to the authentic standard.
(
4) Howitz, K. T.; Bitterman, K. J.; Cohen, H. Y.; Lamming, D. W.; Lavu,
S.; Wood, J. G.; Zipkin, R. E.; Chung, P.; Kisielewski, A.; Zhang, L. L.;
Scherer, B.; Sinclair, D. A. Nature (London) 2003, 425, 191-196.
All the resveratrol produced accumulated inside the cells and was
in the trans aglycone form, as confirmed by LC-MS analysis
-
1
(5) Valenzano, D. R.; Terzibasi, E.; Genade, T.; Cattaneo, A.; Domenici, L.;
Cellerino, A. Curr. Biol. 2006, 16, 296-300.
(
Figure S6). In mammalian cells, 0.34 µg mL (cell volume)
resveratrol was produced. When fed with tyrosine, both 4-coumaric
acid and trans-resveratrol were detected in the cotransfected lines,
indicating that both TAL and the fusion protein are functional
(6) Kaeberlein, M.; McDonagh, T.; Heltweg, B.; Hixon, J.; Westman, E. A.;
Caldwell, S. D.; Napper, A.; Curtis, R.; DiStefano, P. S.; Fields, S.;
Bedalov, A.; Kennedy, B. K. J. Biol. Chem. 2005, 280, 17038-17045.
(7) Kyndt, J. A.; Meyer, T. E.; Cusanovich, M. A.; Van Beeumen, J. J. FEBS
Lett. 2002, 512, 240-244.
(
Figure 2B,E). The production of resveratrol was estimated at 85
(
8) (a) Ralston, L.; Subramanian, S.; Matsuno, M.; Yu, O. Plant Physiol.
-1
ng mL (cell volume). Once again, all the resveratrol accumulated
within the cells. Interestingly, without substrate feeding, the
cotransfected cells synthesized 4-coumaric acid and trans-resvera-
trol, using phenylalanine and/or tyrosine that are present endog-
enously or in the cultured medium (Figure 2C,F). Without substrate
feeding, the cells yielded 28.3 ng mL of resveratrol. Comparison
of the yields from these three experiments indicates that the supply
of phenylalanine or tyrosine is limiting in mammalian cells. Taken
together, introducing the TAL, 4CL, and STS genes into a
mammalian cell line leads to de novo resveratrol biosynthesis.
Our results demonstrate that using unnatural fusion proteins for
metabolic engineering offers a strategy for improving pathway
2
005, 137, 1375-1388. (b) Watts, K. T.; Lee, P. C.; Schmidt-Dannert,
C. ChemBioChem 2004, 5, 500-507. (c) Jiang, H.; Wood, K. V.; Morgan,
J. A. Appl. EnViron. Microbiol. 2005, 71, 2962-2969.
(
9) Becker, J. V.; Armstrong, G. O.; van der Merwe, M. J.; Lambrechts, M.
G.; Vivier, M. A.; Pretorius, I. S. FEMS Yeast Res. 2003, 4, 79-85.
(
10) Romero-Perez, A. I.; Ibern-Gomez, M.; Lamuela-Raventos, R. M.; de La
Torre-Boronat, M. C. J. Agric. Food Chem. 1999, 47, 1533-1536.
-1
(11) (a) Jang, S. K.; Davies, M. V.; Kaufman, R. J.; Wimmer, E. J. Virol.
1
989, 63, 1651-1660. (b) Jang, S. K.; Krausslich, H. G.; Nicklin, M. J.;
Duke, G. M.; Palmenberg, A. C.; Wimmer, E. J. Virol. 1988, 62, 2636-
2643.
(
12) (a) Boshart, M.; Weber, F.; Jahn, G.; Dorsch-Hasler, K.; Fleckenstein,
B.; Schaffner, W. Cell 1985, 41, 521-530. (b) Nelson, J. A.; Reynolds-
Kohler, C.; Smith, B. A. Mol. Cell. Biol. 1987, 7, 4125-9.
13) Stock, P. G.; Bluestone, J. A. Annu. ReV. Med. 2004, 55, 133-56.
(
JA0622094
J. AM. CHEM. SOC.
9
VOL. 128, NO. 40, 2006 13031

 Zhang, Yansheng
Zhang, Yansheng
 Li, Song-Zhe
Li, Song-Zhe
 Li, Jia
Li, Jia
 Pan, Xiangqing
Pan, Xiangqing
 Cahoon, Rebecca E.
Cahoon, Rebecca E.
 Jaworski, Jan G.
Jaworski, Jan G.
 Wang, Xuemin
Wang, Xuemin
 Jez, Joseph M.
Jez, Joseph M.
 Chen, Feng
Chen, Feng
 Yu, Oliver
Yu, Oliver