JOURNAL OF THE CHINESE
CHEMICAL SOCIETY
2
,3-Butanediol Dehydrogenase
extract, 2% peptone, 2% glucose) containing 0.36 mM acetoin.
The production of active recombinant TcBdh was shown by the
enzyme assay.
RESULTS AND DISCUSSION
Cloning, characterization of a cDNA encoding TcBdh
and a 3-D structural model of TcBdh
Expression and purification of the recombinant TcBdh:
The transformed yeast containing the TcBdh gene was grown at
A putative TcBdh cDNA clone was identified based
on its sequence homology to the published Bdhs in NCBI
website. The coding region of TcBdh cDNA was 1224 bp
that encodes a protein of 408 amino acid residues with a
calculated molecular mass of 49.3 kDa (GenBank acces-
sion JF896462). Theoretical pI/Mw is 6.15/49300. A
homology search using DELTA-BLAST detected putative
conserved domains of medium chain dehydrogenase/re-
ductase (MDR) superfamily. The conserved domains in-
3
2
4
0 °C in 250 mL of YPD medium containing 0.36 mM acetoin for
days. The cells were harvested and soluble proteins extracted in
0 mM Tris-HCl, pH 7.0, containing with glass beads as de-
1
scribed before. The recombinant TcBdh was purified by Ni-NTA
6
affinity chromatography (elution buffer: 40 mM Tris-HCl, pH
7
.0, containing 5-250 mM imidazole) according to the manufac-
ture’s instruction (Qiagen). The purified protein was checked by
2% SDS-PAGE. The purified protein was pooled and centri-
2
+
38
40
60
cluded a catalytic Zn binding site at Cys , Ser , His ,
1
1
68
90
93
96
Asp , a structural Zn binding site at Cys , Cys , Cys ,
fuged to remove salt using Amicon membrane (5000 MW), fi-
nally the recombinant TcBdh (0.16 µg/µL) was in 20 mM
Tris-HCl containing 2.5 mM imidazole, 45% glycerol. Proteins
on gel were detected by staining with Coomassie Brilliant Blue
R-250. Protein concentration was determined by a Protein Assay
Kit (Bio-Rad, Richmond, CA) using bovine serum albumin as a
standard.
1
Cys , and a NAD binding site at Cys Gly Ser , His ,
04
+
38
39
40
43
8
6
168
172
191
192
194
195
Phe , Asp , Thr , Trp Gly , Gly Pro Ile
196
,
,
2
15
216
217
221
236
260
261
Ile Asp Arg , Arg , Phe , Cys Gly Thr
262
2
86
303
304
305
329
330
331
Ile , Ile Ala Val , Gly Gln Ala , Phe . Fig.
372
1
A shows the optimal alignment of the amino acid se-
quences of TcBdh with 4 related Bdh sequences from other
sources. The TcBdh shares a 68% identity with R,R-bu-
tanediol dehydrogenase of ScBdh (Saccharomyces cerevisiae
S288c, NP_009341), a 67% with R,R-butanediol dehydro-
genase of CnBdh (Cryptococcus neoformans var. neo-
formans JEC21, XP_568483), a 66% with R,R-butanediol
dehydrogenase of CgBdh (Cryptococcus gattii WM276,
accession no. XP_003197328), and a 60% identity with
CnBdh (Cryptococcus neoformans var. neoformans JEC21,
XP_568483). The N-terminal end of the protein also con-
tains GroES-like domain as identified by InterProScan
Activity assay and kinetic studies of the recombinant
TcBdh: Bdh activity was determined by measuring NADH oxida-
2
1
tion. The reaction mixture (100 mL) contained 33 mM potas-
sium phosphate, pH 7.0, 0.2 mM NADH, 25 mM acetoin. The re-
action was started by the addition of 2 mg (2 nM) TcBdh. The reac-
tion was followed by the decrease in A340 due to the oxidation of
NADH.
The kinetic properties of the TcBdh (2 mg) was determined
by varying the concentrations of acetoin (7 to 25 mM) with fixed
amount of 0.2 mM NADH. The change in absorbance at 340 nm
was recorded between 10 sec and 40 sec. The molar absorption
(
http://www.ebi.ac.uk/Tools/pfa/iprscan/). The secondary
structure (Fig. 1A, represented as a helices and b strands)
and a 3-D structural model (Fig. 1B, represented as solid
ribbon) were predicted using SWISS-MODEL program.
The 3-D structural model (Fig. 1B, light blue) was con-
structed based on the known crystal structure of Pseudo-
monas putida formaldehyde dehydrogenase (PpFdh, PDB
-
1
-1
coefficient of NADH at 340 nm is 6.22 mM cm . The K
M
, Vmax
and kcat were calculated from Lineweaver-Burk plots.
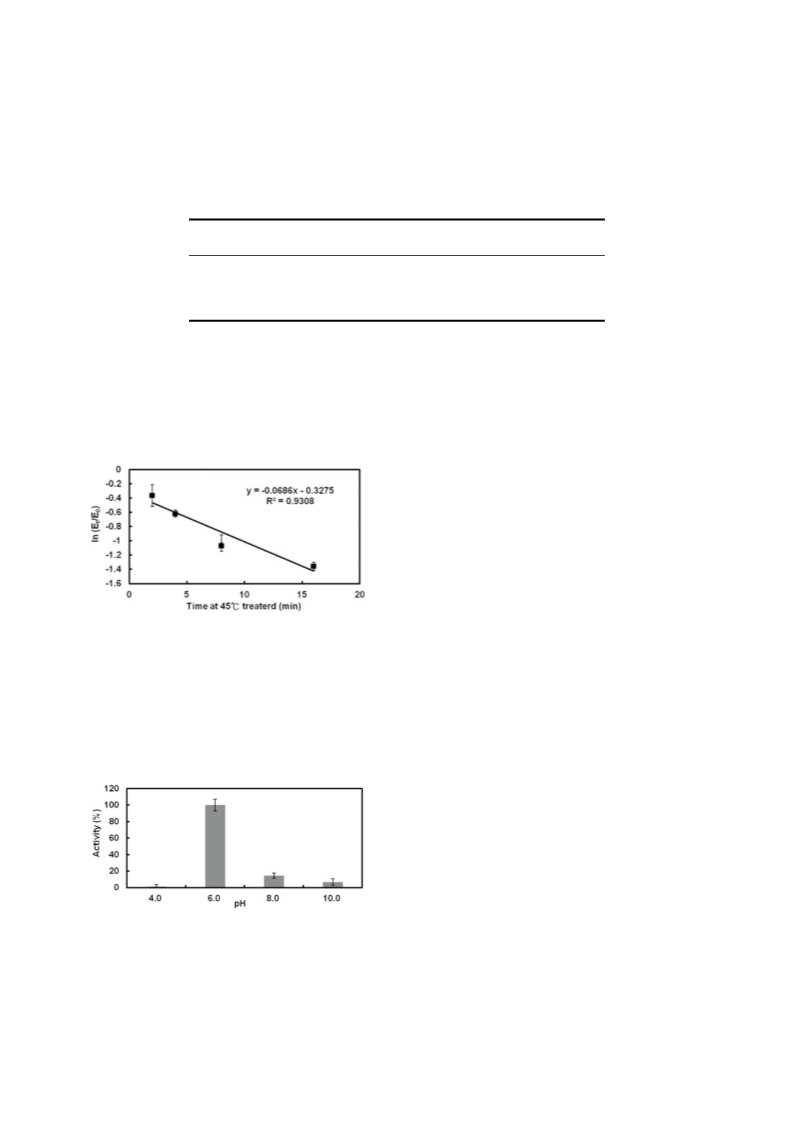
Biochemical properties: The stability of TcBdh under var-
ious conditions was studied by assaying its ability to reduce
acetoin as described above. Aliquots of the TcBdh sample were
tested for: (1) Thermal effect. Enzyme sample (2 mg/24 mL en-
zyme in 20 mM Tris-buffer, pH 7.0, containing 5% glycerol per
reaction) was heated to 45 °C for 2, 4, 8 or 16 min. (2) pH effect.
Enzyme sample (2 mg/24 mL enzyme in 20 mM Tris-buffer, pH
192
194
197
ID: 1KOL, white). Red box (Fig. 1A, Gly XGly XXGly )
indicates the highly conserved Gly-X-Gly-X-X-Gly se-
quence found in the MDR (medium chain dehydrogenase/
2
2
reductase, Nordling et al. 2002 ) family and residues
7
.0, containing 5% glycerol per reaction) was adjusted to desired
found in the coenzyme-binding pocket. Red denotes struc-
+
ture of NAD (Fig. 1B).
pH by adding a half volume of buffer with different pHs: 0.2 M ci-
trate buffer (pH 4.0), 0.2 M potassium phosphate buffer (pH 6.0,
or 8.0) or 0.2 M CAPS buffer (pH 10.0). Each sample was incu-
Expression and purification of the recombinant
TcBdh
o
bated at 37 C for 30 min. After each treatment, the residual Bdh
The coding region of TcBdh (1,224 bp) was amplified
by PCR and subcloned into a yeast expression vector,
activity was tested as described above.
J. Chin. Chem. Soc. 2015, 62, 443-448
© 2015 The Chemical Society Located in Taipei & Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim
www.jccs.wiley-vch.de
445

 Ken, Chuian-Fu
Ken, Chuian-Fu
 Tsai, Wei-Wei
Tsai, Wei-Wei
 Wen, Lisa
Wen, Lisa
 Sheu, Dey-Chyi
Sheu, Dey-Chyi
 Lin, Chi-Tsai
Lin, Chi-Tsai