676
Alterio et al.
tration of 1 mM were carried out at 208C in 20 mM sodium phos-
phate buffer pH 7.4. The hydrolysis of S-derivatives of glutathione
was determined by monitoring the respective decrease in absorb-
ance at 240 nm.
3. Lee, E. Y.; Lee, W. H. Proc Natl Acad Sci USA 1986, 83, 6337–
6341.
4. Degrassi, G.; Uotila, L.; Klima, R.; Venturi, V. Appl Environ
Microbiol 1999, 65, 3470–3472.
5. Yurimoto, H.; Lee, B.; Yano, T.; Sakai, Y.; Kato, N. Microbiology
2003, 149, 1971–1979.
Crystallization and X-Ray Data Collection
6. Herring, C. D.; Blattner, F. R. J Bacteriol 2004, 186, 6714–6720.
7. Gonzales, C. F.; Proudfoot, M.; Brown, G.; Korniyenko, Y.;
Mori, H.; Savchenko, A. V.; Yakunin, A. F. J Biol Chem 2006,
281, 14514–14522.
PhEst was crystallized at 293 K using the hanging drop vapor diffu-
sion technique. Drops were prepared by mixing 1 ll of enzyme solu-
tion (4 mg ml21 in 25 mM TRIS-HCl, pH 7.3) with 1 ll of precipi-
tant solution (20% (w/v) PEG MME 5000, 0.2M sodium acetate,
0.1M TRIS-HCl, pH 7.0), and equilibrated over a well containing 1
ml of precipitant solution. Crystals appeared after 3 days and grew
in about one week to maximum dimensions of 0.3 3 0.2 3 0.2
8. Kordic, S.; Cummins, I.; Edwards, R. Arch Biochem Biophys
2002, 399, 232–238.
9. Haslam, R.; Rust, S.; Pallett, K.; Cole, D.; Coleman, J. Plant
Physiol Biochem 2002, 40, 281–288.
3
˚
mm . A complete dataset was collected at 2.20-A resolution from a
10. Cummins, I.; McAuley, K. M.; Fordham-Skelton, A.; Schwoerer,
R.; Steel, P. G.; Davis, B. G.; Edwards, R. J Mol Biol 2006, 359,
422–432.
single crystal at the temperature of 100 K, with a copper rotating an-
ode generator developed by Rigaku and equipped with Rigaku Sat-
urn CCD detector. Prior to cryogenic freezing, crystals were trans-
ferred to the precipitant solution with the addition of 15% (w/v)
glycerol. Data were processed using the HKL2000 crystallographic
data resolution package (Denzo/Scalepack).47 The crystals belonged
to the space group P212121 with unit cell dimensions of a 5 49.49
11. Legler, P. M.; Kumaran, D.; Swaminathan, S.; Studier, F. W.;
Millard, C. B. Biochemistry 2008, 47, 9592–9601.
12. Wu, D.; Li, Y.; Song, G.; Zhang, D.; Shaw, N.; Liu, Z. J. FASEB J
2009, 23, 1441–1446.
13. van Straaten, K. E.; Gonzalez, C. F.; Valladares, R. B.; Xu, X.; Sav-
chenko, A. V.; Sanders, D. A. Protein Sci 2009, 18, 2196–2202.
14. Aurilia, V.; Parracino, A.; Saviano, M.; Rossi, M.; D’Auria, S.
Gene 2007, 397, 51–57.
˚
˚
˚
A, b 5 129.75 A, c 5 152.67 A. The Matthews coefficient (VM
5
1.99 A Da21) indicated that the crystallographic asymmetric unit
contained four molecules according to a solvent content of 38%.
Data collection statistics are reported in Table I.
3
˚
15. Laskowsky, R. A.; MacArthur, M. W.; Moss, D. S.; Thornton, J.
M. J Appl Cryst 1993, 26, 283–291.
16. Hutchinson, E. G.; Thornton, J. M. Protein Sci 1996, 5, 212–
220.
Structure Determination and Refinement
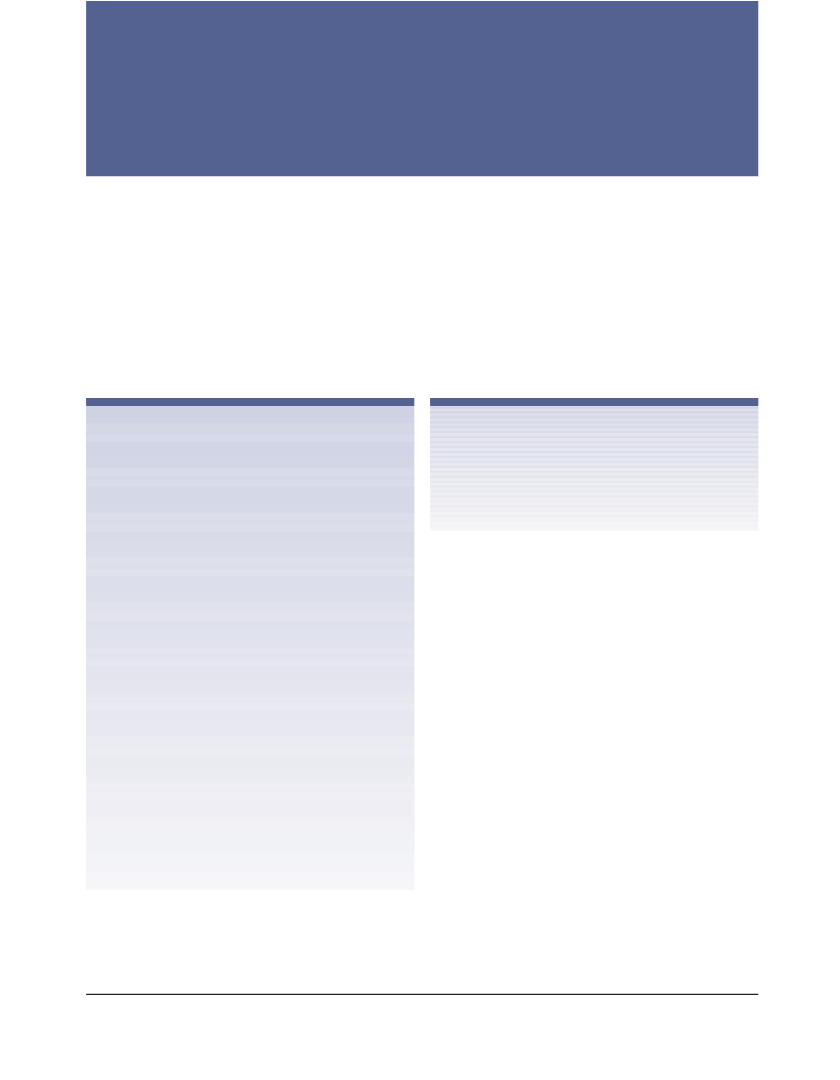
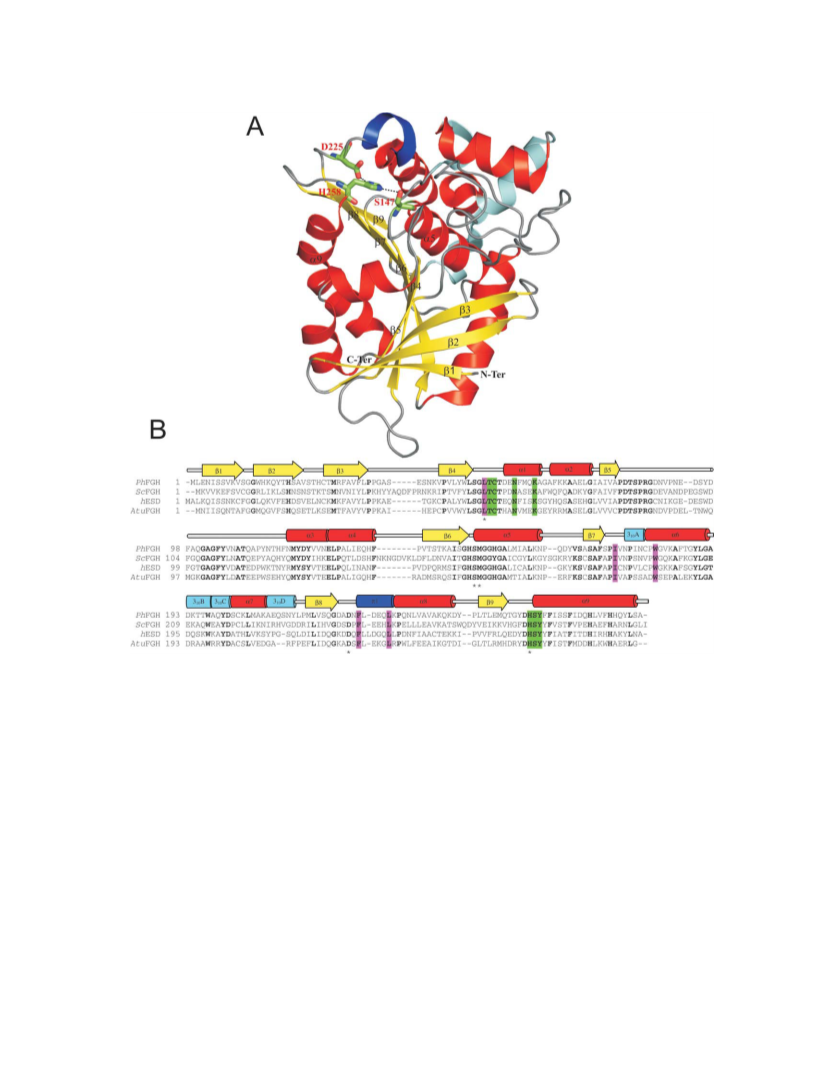
The structure of PhEst was solved by molecular replacement tech-
nique using the program AMoRe48 and the crystallographic struc-
ture of the ScFGH (PDB code 1PV1)11 as model template. The rota-
tion and translation functions were calculated using data between
17. De Simone, G.; Galdiero, S.; Manco, G.; Lang, D.; Rossi, M.;
Pedone, C. J Mol Biol 2000, 303, 761–771.
18. De Simone, G.; Mandrich, L.; Menchise, V.; Giordano, V.; Feb-
braio, F.; Rossi, M.; Pedone, C.; Manco, G. J Biol Chem 2004,
279, 6815–6823.
˚
15.0 and 3.5 A resolution. The one-body translation search, using
the centred-overlap function (c-o), on the first 50 rotation solutions
led to a single solution with a correlation coefficient of 25.4% and
an R-factor of 52.1%. The n-body translation search carried out
with the phased-translation function (p-t), by including a PC
refinement before each n-body translation search led to finding the
remaining three molecules contained into the asymmetric unit. This
improved the correlation coefficient and the R-factor to 73.8 and
32.7%, respectively. Refinement of the structure was carried out
using CNS23 and model building was performed with O.49 The first
19. Bentahir, M.; Feller, G.; Aittaleb, M.; Lamotte-Brasseur, J.;
Himri, T.; Chessa, J. P.; Gerday, C. J Biol Chem 2000, 275,
11147–11153.
20. Hoyoux, A.; Jennes, I.; Dubois, P.; Genicot, S.; Dubail, F.; Fran-
c¸ois, J. M.; Baise, E.; Feller, G.; Gerday, C. Appl Environ Micro-
biol 2001, 67, 1529–1535.
21. Lonhienne, T.; Zoidakis, J.; Vorgias, C. E.; Feller, G.; Gerday, C.;
Bouriotis, V. J Mol Biol 2001, 310, 291–297.
22. Tina, K. G.; Bhadra, R.; Srinivasan, N. Nucleic Acids Res 2007,
35, W473–W476.
cycles of the refinement were carried out with four-fold NCS-
restraints with an energy barrier of 300 kcal mol21 A . After R-fac-
2
˚
¨
23. Brunger, A. T.; Adams, P. D.; Clore, G. M.; De Lano, W. L.;
tor and R-free reached 21.7 and 24.1%, respectively, the NCS
restraints were removed, and further cycles of manual rebuilding
and positional and temperature factor refinement were necessary to
reduce the crystallographic R-factor and R-free values (in the 20.00-
Gros, P.; Grosse-Kunstleve, R. W.; Jiang, J. S.; Kuszewski, J.;
Nilges, M.; Pannu, N. S.; Read, R. J.; Rice, L. M.; Simonson, T.;
Warren, G. L. Acta Crystallogr Sect D 1998, 54, 905–921.
24. Nicholls, A.; Sharp, K. A.; Honig, B. Proteins 1991, 11, 281–296.
25. Feller, G.; Arpigny, J. L.; Nminx, E.; Geday, C. Camp Biochem
Physiol 1997, 118A, 495–499.
˚
to 2.20-A resolution range) to 16.1 and 20.5%, respectively. Data
refinement statistics are summarized in Table I. Coordinates and
structure factors were deposited in the Protein Data Bank (accession
code 3LS2).
26. Siddiqui, K. S.; Cavicchioli, R. Annu Rev Biochem 2006, 75,
403–433.
27. Saunders, N. F.; Thomas, T.; Curmi, P. M.; Mattick, J. S.; Kuc-
zek, E.; Slade, R.; Davis, J.; Franzmann, P. D.; Boone, D.; Ruster-
holtz, K.; Feldman, R.; Gates, C.; Bench, S.; Sowers, K.; Kadner,
K.; Aerts, A.; Dehal, P.; Detter, C.; Glavina, T.; Lucas, S.;
Richardson, P.; Larimer, F.; Hauser, L.; Land, M.; Cavicchioli, R.
Genome Res 2003, 13, 1580–1588.
REFERENCES
1. Harms, N.; Ras, J.; Reijnders, W. N.; van Spanning, R. J.; Stout-
hamer, A. H. J Bacteriol 1996, 178, 6296–6299.
2. Uotila, L.; Koivusalo, M. J Biol Chem 1974, 249, 7664–7672.
Biopolymers

 Alterio, Vincenzo
Alterio, Vincenzo
 Aurilia, Vincenzo
Aurilia, Vincenzo
 Romanelli, Alessandra
Romanelli, Alessandra
 Parracino, Antonietta
Parracino, Antonietta
 Saviano, Michele
Saviano, Michele
 D'Auria, Sabato
D'Auria, Sabato
 de Simone, Giuseppina
de Simone, Giuseppina